- ... all
![[*]](/icons/latex2html/icons/footnote.png)
- http://www2.ebi.ac.uk/pdb/
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... resource
![[*]](/icons/latex2html/icons/footnote.png)
- http://scop.mrc-lmb.cam.ac.uk/scop/
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... CATH
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/bsm/cath/
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
site
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/bsm/cath/lex/cathinfo.html
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... correct?
![[*]](/icons/latex2html/icons/footnote.png)
- Although, interestingly, 75% is
actually three-quarters of the way between the random alignment
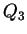 of
35% and 88%
of
35% and 88%
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
Martin
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/~martin/PROGRAMS.TXT
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...ProFit
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/~martin/#programs
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...)
![[*]](/icons/latex2html/icons/footnote.png)
- this is the solvent accessible contact surface area buried
on complexing with antigen
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... class
![[*]](/icons/latex2html/icons/footnote.png)
- based on a simple
clustering of interface area
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
topography
![[*]](/icons/latex2html/icons/footnote.png)
- combining site topography classification
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... TSA
![[*]](/icons/latex2html/icons/footnote.png)
- transition state analogue
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... topography
![[*]](/icons/latex2html/icons/footnote.png)
- combining site topography classification
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
internet
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/~martin/abs/GeneralInfo.html#contactdata
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... GRASP
![[*]](/icons/latex2html/icons/footnote.png)
- GRASP macros are available from
http://www.biochem.ucl.ac.uk/~bob/Ab/graspMacro.txt
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... alignments
![[*]](/icons/latex2html/icons/footnote.png)
- often generated automatically
using substitution
matrices
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
series
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.mbi.ucla.edu/people/fischer/BENCH/benchmark1.html
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...)
![[*]](/icons/latex2html/icons/footnote.png)
- This manipulation of the results is not completely
valid. Ideally, the whole experiment should be repeated without any
highly similar pairs, rather than just discarding their results as we
have done here.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ... SOM\_PAK
![[*]](/icons/latex2html/icons/footnote.png)
- source code available from
http://nucleus.hut.fi/nnrc/nnrc-programs.html
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
trend
![[*]](/icons/latex2html/icons/footnote.png)
- We need to verify that this is not an effect of sequence
order, by mapping reversed or shuffled sequences. The order of training
vector presentation should not affect the mapping.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
internet
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/~bob/thesis/1sxaA0_anim.html
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
- ...
Web
![[*]](/icons/latex2html/icons/footnote.png)
- http://www.biochem.ucl.ac.uk/bsm/cath/
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.